Posts
My thoughts and ideas
Welcome to the blog
My thoughts and ideas
Introduction to bioinformatics for RNA sequence analysis
We offer several options for completing the hands-on components of this course including Amazon Cloud (AWS), Google Cloud (GCP), and Docker images.
Using cloud computing (AWS or GCP) to complete this course involves two major components: (1) Launching an instance on the cloud (essentially renting a virtual computer by the hour and turning it on) and (2) logging into that instance.
Using Docker involves running a precompiled image locally on your computer. For specific instructions on how to use each, choose one of the options below:
Covered in this section: logging into AWS EC2 console, starting an instance from the course AMI and configuring it in the console (select instance AMI, instance type, instance details, storage volumes, tags, security group, and key pairs).
In the previous section Introduction to AWS we reviewed fundamental concepts of cloud computing and some of the jargon and features specific to AWS. In this section we will learn how to launch an instance specifically for this course.
In order to launch your own instance you will either need to use your own personal AWS account, an account provided through your institution OR if you are taking a live version of this course you will be assigned an AWS account using the IAM system. The following instructions are used for a live version of this course, but the process would be essentially the same when using a personal or institutional account.
Briefly the process for launching an EC2 instance for this course involves these steps:
chmod 400 cshl_2024_student.pem
ssh -i cshl_2024_student.pem ubuntu@[your ip address or dns name] # [your ip address or dns name] is the public IP address of your instance
-i selects a file from which the public key authentication is read. ubuntu is the name of a user on the system you are logging into (a default user of the Ubuntu operating system). [your ip address] is the address of the linux system on Amazon that you are logging into. Instead of ip address you can also use a public dns name.
Open a PowerShell session , and follow the steps as listed in Logging in to AWS with Terminal.
To configure PuTTy, start PuTTy and do the following:
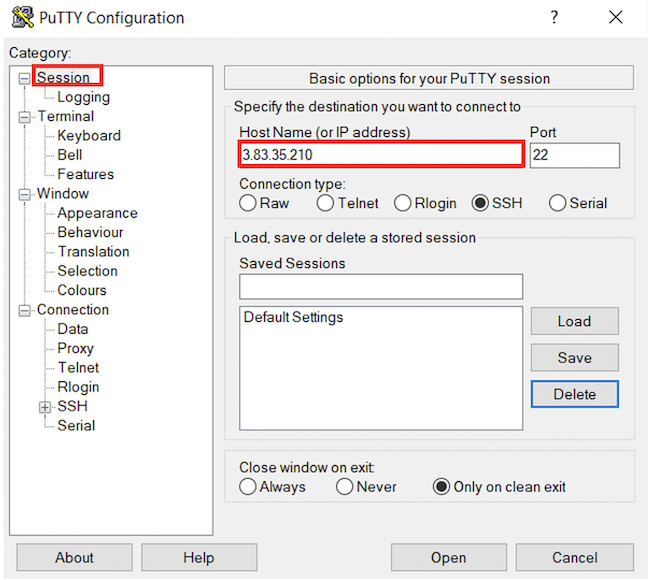
ubuntu.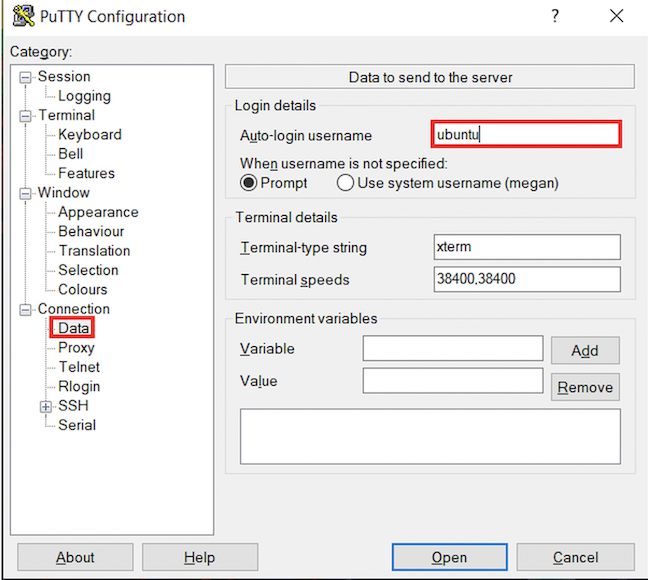
cshl_2024_student.ppk certificate that you downloaded above.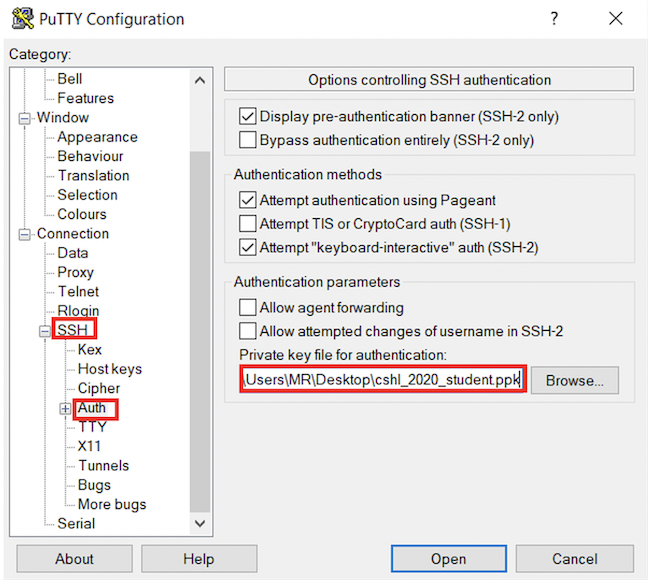
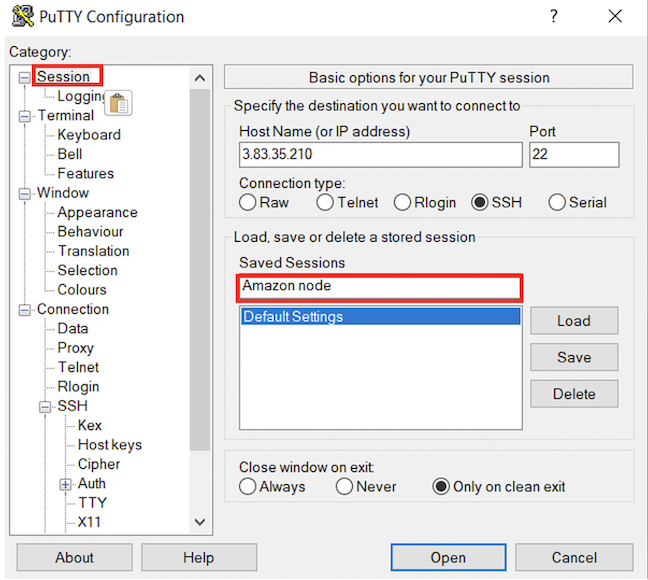
scp -i cshl_2024_student.pem ubuntu@[your ip address or dns name]:nice_alignments.bam .
http://[your ip address]/ or http://[your dns name]
When you log in, you will notice that you have one directory already: “workspace”.
If you would like to upload your data to the AWS instance, use the example scp command below. Be sure to replace the variables below with the local path to your data, MY_DATA, and the amazon instance IP, YOUR_IP_ADDRESS.
scp -i cshl_2024_student.pem __MY_DATA__ ubuntu@[your ip address or dns name]:/
If you are trying to do this course on your own using the online materials only, of course an AWS EC2 instance has not been set up for you. If you have access to an AWS account though you can can start with the same Amazon AMI we use to create instances for each student. Currently this is:
Name: cshl-seqtec-2024 available in the US East, N. Virginia region (us-east-1).
We typically use an instance type of m6a.xlarge. For detailed instructions on how we created the AMI and configure each instance, please refer to the AWS Setup page.
Once you are logged into your EC2 instance using ssh, the following commands can be used to explore basic information about the machine
#How are storage volumes mounted?
lsblk
#How much storage space in being used for various mount points?
df -h
#Detailed description of hardware
lshw
#See active resource usage right now with "top". Use "1" to see all CPUs individually. Use "q" to exit
top
Coming Soon
We have built a docker image with all necessary software tools installed to complete this course. This allows the hands-on exercises to be perfomed on your own desktop or laptop or anywhere that has docker installed. The following commands can be used to download and run this image as an interactive docker container.
First you will need to install docker. Typically this involves installing Docker Desktop. Once installed, start docker, and open a terminal application. Explanations of how to do this are detailed above in the Using AWS section. At the terminal you can test your docker installation is working/running with docker --help. Once you have docker working you can use the following commands to download, set up, and run the rnabio image.
docker pull griffithlab/rnabio:0.0.3
mkdir -p ~/rnabio-workspace
docker run -v ~/rnabio-workspace:/workspace:rw --user ubuntu:ubuntu -it griffithlab/rnabio:0.0.3 /bin/bash
The docker session should now be ready for the Unix tutorial and practical exercises.